Research
Research topics
Interactions between biomolecules are formed by specific multipoint binding through a combination of non-covalent bonds. In order to clarify the contribution to these interactions, we are analyzing various proteins, including protein-protein interactions in general, antibody-antigen, receptor-ligand, protein-metal ion, and multimeric proteins. We are attempting to elucidate the mechanism of specificity and affinity creation by precisely analyzing and discussing biomolecular interactions from multiple angles using kinetic/thermodynamic analysis using physical chemistry methods such as isothermal titration calorimetry (ITC), surface plasmon resonance (SPR), differential scanning calorimetry (DSC), differential scanning fluorescence (DSF), and microscale thermophoresis (MST), spectroscopic analysis including circular dichroism (CD), as well as mass spectrometry and crystal structure analysis. The obtained physicochemical parameters are used for the design of proteins and small molecules for medical and material applications by integrating in vitro and in silico approaches with computational and information science to incorporate the viewpoints of molecular dynamics and ensemble. Based on these findings, we aim to develop functional molecules with high functionality and excellent physical properties, and to explore molecular species that control specific molecular interactions, which will lead to the construction of a basis for drug discovery and the understanding of the nature of “interactions” in life.
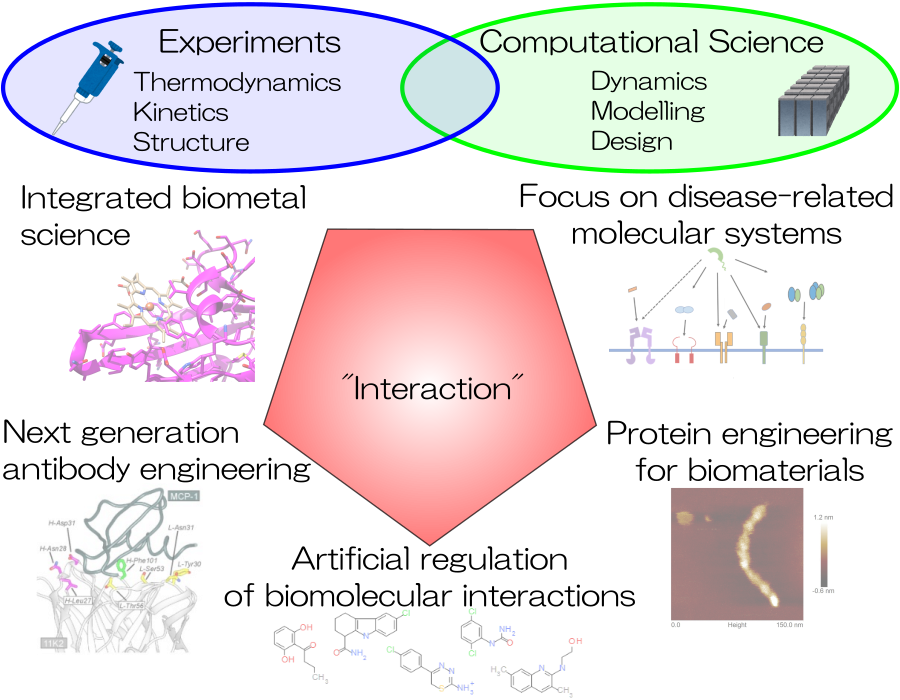
Research theme
- Next-Generation Antibody Engineering
- Molecular Machinery Elucidation of Disease-Related Proteins
- Bio-metal Science
- Development of Bio-molecular Interaction Control Agents
- Protein Engineering for Biomaterials
- Computational Chemistry and Biology for Molecular Design
Next-Generation Antibody Engineering:
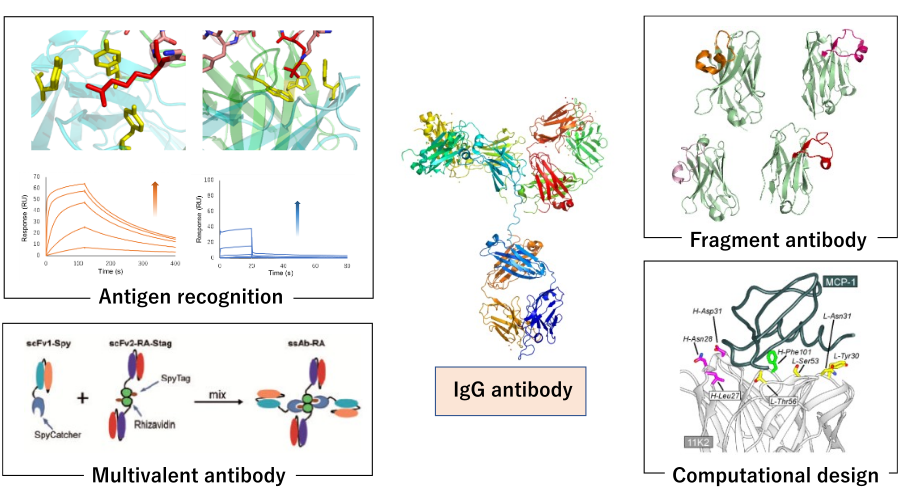
Antibodies are widely used as biopharmaceuticals and diagnostics due to their ability to recognize antigen molecules with high affinity and specificity. In recent years, the biopharmaceutical market centered on antibody drugs has continued to expand, and next-generation antibodies are needed to further improve their properties and functionalities. In our laboratory, we are conducting research that contributes to the creation of next-generation antibody molecules by analyzing the antigen recognition mechanisms of antibodies from multiple perspectives and by utilizing protein engineering techniques. In particular, we have focused on research that efficiently performs antibody design and affinity enhancement, which were previously difficult, by combining computational science and experimental techniques.
Specifically, we seek a detailed understanding at the molecular level by combining physical and chemical analyses of antibody properties such as SPR, ITC and CD, including antigen recognition, thermal stability and aggregation, with structural analysis by X-ray crystallography and cell biological methods. We also work on antibody modification using evolutionary engineering techniques such as phage display and cDNA display, single domain antibody (VHH) development, antibody miniaturization, fusion antibody production, and rational design of antibody functional modification using computational science. Recently, we have also focused on research that incorporates high-throughput screening technologies and data-driven design to accelerate antibody design and evaluation.
Projects
- Elucidation of the molecular recognition mechanisms of antibodies: specific antibodies for post-translational modifications from protein antigens, etc.
- Functional enhancement through antibody engineering combining biochemical and structural biology methods
- Establishment of a foundation for antibody development that integrates high-throughput screening technology and data-driven design
- Development of new antibody formats based on diverse molecular recognition mechanisms
- Development of antibody design techniques based on computational science (molecular dynamics calculations, free energy calculations, etc.)
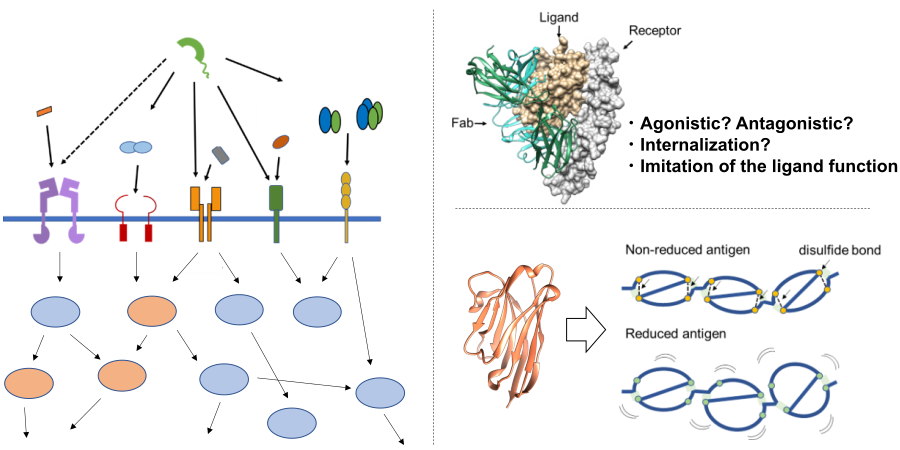
Molecular Machinery Elucidation of Disease-Related Proteins:
Proteins build complex life systems by interacting with various biomolecules in living organisms, and the breakdown of this tightly controlled system leads to various diseases such as cancer, autoimmune diseases and neurodegenerative diseases. For example, various tumor suppressor proteins in the body play a role in maintaining cellular homeostasis, but a decrease in their expression or mutations in their sequences can disrupt their functions, leading to carcinogenesis. In addition, various proteins interact not only within individual organisms, but also between pathogenic microorganisms that cause infections and their hosts, and in the case of the novel coronavirus, which has recently become a major public concern, the interaction between receptors expressed on human cell surfaces and viral surface proteins plays a crucial role. In our laboratory, we focus on groups of proteins associated with various diseases and aim to elucidate the mechanisms of these diseases at the molecular level by unraveling the molecular machinery that links them. We analyze the molecular functions of disease-associated proteins from different angles using various methods such as physical and chemical analyses, structural analyses, cell biological analyses, and molecular dynamics simulations, and based on the insights gained, we conduct research aimed at controlling protein function using various modalities such as antibodies, mid-sized molecules, and small molecules.
Projects
- Analysis and molecular modification of the molecular function mechanisms of tumor suppressor proteins
- Elucidation of the degradation mechanisms of tumor suppressor proteins and inhibitor discovery
- Structural and functional analysis of synapse formation-related proteins and development of regulatory molecules
- Molecular mechanism analysis of extracellular matrix and cell adhesion factors and development of functional regulatory molecules
- Analysis of antigen recognition mechanisms by autoantibodies in autoimmune diseases
- Functional analysis of surface proteins of pathogenic microbial proteins and inhibitor discovery
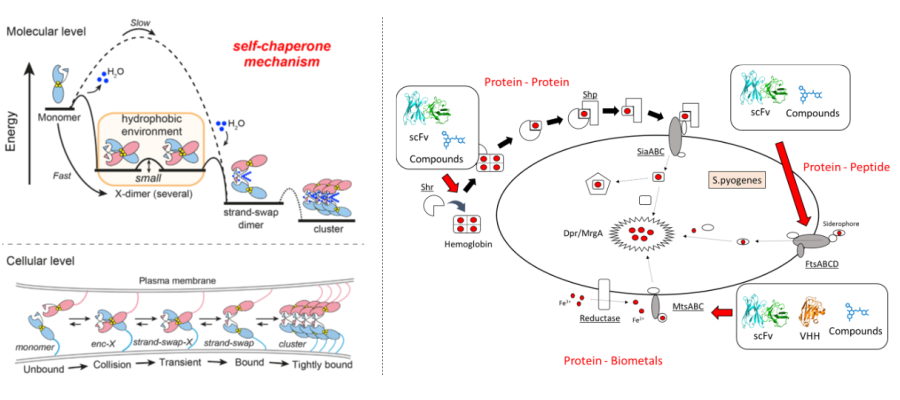
Bio-metal Science:
Metals are essential elements for protein function, and their interactions form the basis of life phenomena. Proteins possessed by a wide variety of organisms, from humans to plants and microorganisms, require metals for their functional expression, and perturbations in this regulation are known to lead to disease. For example, abnormalities in the regulation of metal-dependent proteins can promote the development and metastasis of cancer, and normalizing this regulation is expected to lead to treatments. Furthermore, elucidating the sophisticated mechanisms by which pathogenic microorganisms acquire metals from their hosts and control them is expected to lead to the development of new antibiotics with novel mechanisms of action. In other words, elucidating the relationship between protein function and metals in life deepens our understanding of “life” and leads to the development of treatments for various diseases. In our laboratory, we focus on metal-mediated intercellular adhesion, metal transport, and metal uptake mechanisms of pathogenic microorganisms, and conduct research aimed at elucidating and controlling metal-mediated protein functions. In particular, we focus on elucidating the mechanisms of complex metal-protein interactions at the atomic level by combining various methods such as X-ray crystallography, mass spectrometry, and molecular dynamics simulations.
Projects
- Elucidation and control of the assembly mechanisms of metal-dependent cell adhesion proteins
- Elucidation of metal acquisition mechanisms in pathogenic microorganisms and inhibitor development
- Elucidation of molecular recognition mechanisms of metal-binding proteins and development of selectively targeting antibodies
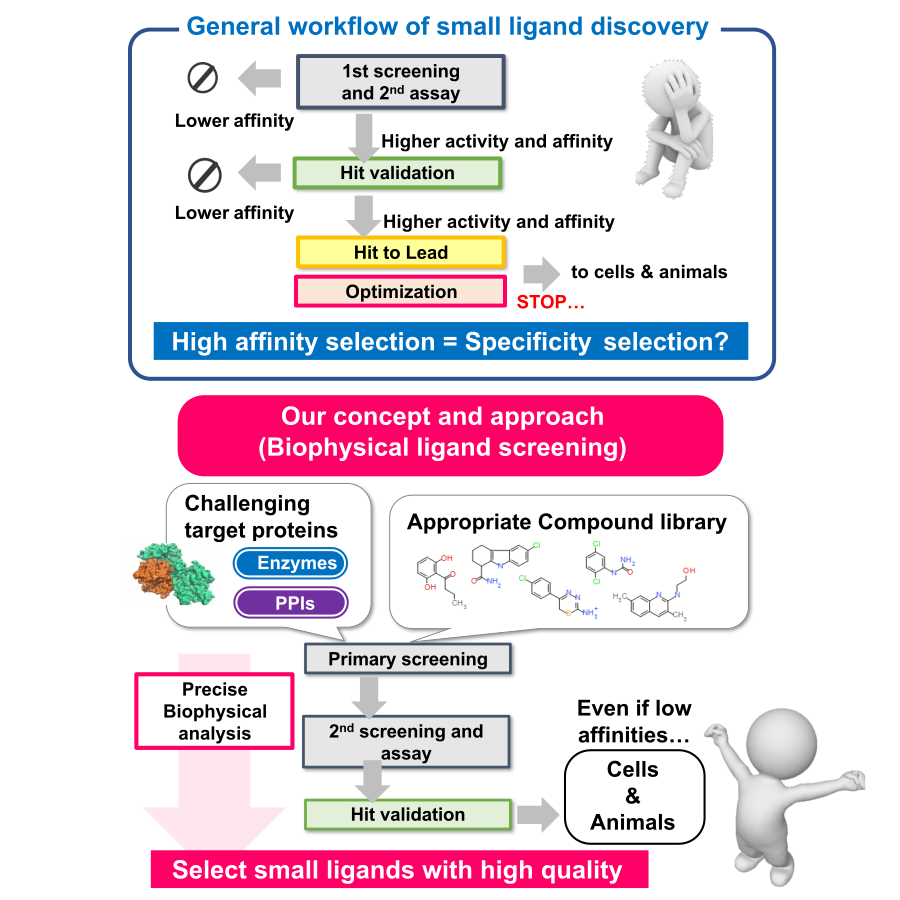
Development of Bio-molecular Interaction Control Agents:
We are working to create small molecule ligands that target the interactions of proteins involved in intracellular signaling and metabolic regulation. In recent drug discovery research, the proportion of targets that are difficult to address with conventional drug discovery techniques, such as membrane proteins, protein-protein interactions (PPIs), and intrinsically disordered proteins, has increased. Therefore, in this laboratory, we conduct research to explore new ligands for challenging targets by accurately evaluating the properties of biomolecules using various physical and chemical analysis techniques (ITC, SPR, DSC, MST, HDX-MS, etc.) and quantitatively analyzing the “quality” of their interactions. Using various compound libraries such as fragment libraries, PPI libraries and natural product libraries, we aim to create functional small molecule ligands that contribute not only to drug discovery but also to life science research, in close collaboration with Drug Discovery Initiative, The University of Tokyo and drug discovery support project of AMED.
Projects
- Development of techniques to analyze interactions between target molecules and low-molecular-weight compounds
- Rational design and optimization of PPI inhibitors
- Exploration of low-molecular-weight compounds for intrinsically disordered proteins
- Design of new functional molecules based on peptide scaffolds
Protein Engineering for Biomaterials:
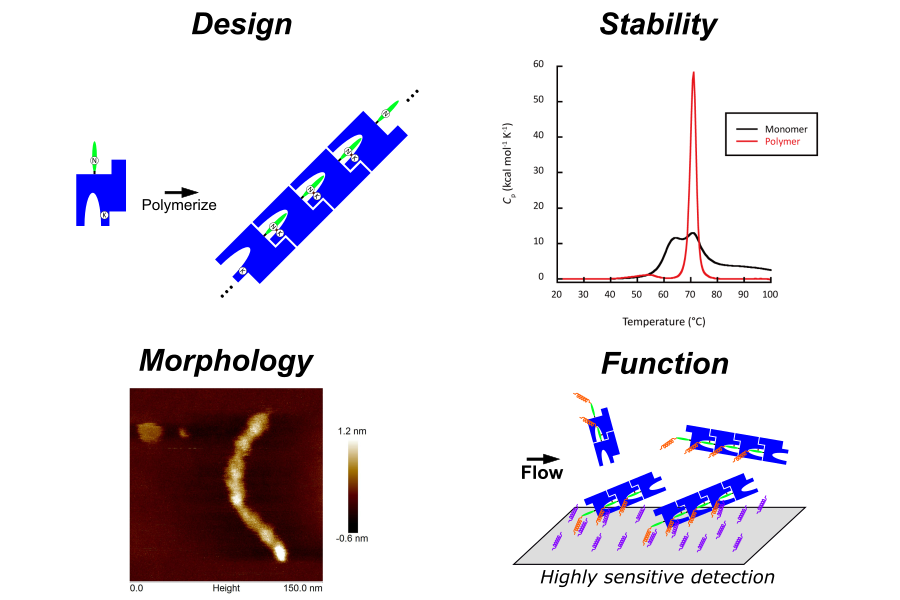
Proteins are fundamental building blocks that support life activities and possess unique properties not found in other substances, such as advanced self-assembly and specific molecular recognition capabilities. Therefore, applications are expected not only in the pharmaceutical field, but also in a wide range of fields such as energy, environment and advanced materials. In our laboratory, we focus on the diverse functions and excellent properties of proteins that make up living organisms, and aim to develop innovative biomaterials with new functions and performances not found in conventional materials by precisely redesigning their amino acid sequences.
Projects
- Supercharging design of antibodies and application to detection materials
- Design of covalent binders utilizing intramolecular/intermolecular cross-linking of microbial-derived adhesive proteins
- Development of ligand proteins used for antibody analysis columns
Computational Chemistry and Biology for Molecular Design:
In relation to the above research themes, we are actively promoting the analysis of biomolecules based on computational and information sciences using the supercomputer at the Institute of Medical Science. Using computers, we can predict the three-dimensional structures and dynamic behaviors of proteins and complexes, and quantitatively evaluate the energies of molecular interactions. In addition, large-scale mutant analyses that are difficult to perform experimentally can be performed efficiently. For example, we can comprehensively analyze the properties of proteins on the computer when each amino acid residue of a given protein is changed to a different amino acid. We are also working to elucidate detailed mechanisms at the molecular level that are difficult to clarify experimentally, using molecular dynamics simulations and free energy calculations. By advancing experiments based on these computational results, we aim to gain a more rational understanding of the nature of “interactions” in life and link it to the development of new biopharmaceuticals and materials creation.




